Open a .csv file for the World maps panel to display a table with their sequences and number of mismatches on each primer
You can search species name as well as sequences on the search box, and download the full table
A high number of mismatches can lead to decrease chances of amplification
Welcome to GAPeDNA
GAPeDNA lets you explore taxonomic coverage gaps in eDNA metabarcoding reference databases. Select a taxon group, spatial resolution, and primer pair to visualise the proportion of species represented in the reference database — and download species lists with their sequences.
How it works
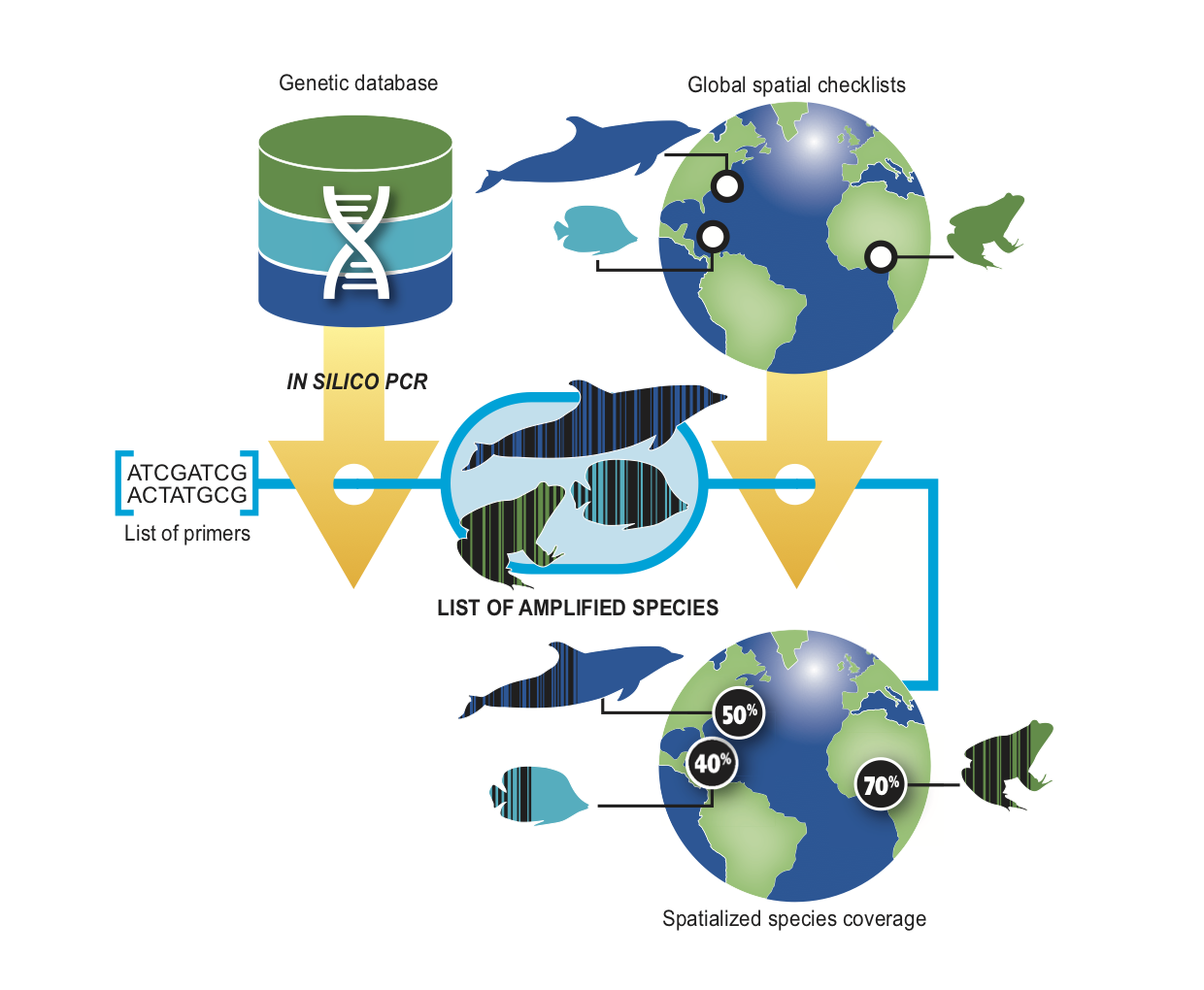
Occurrence data sources
Species-presence matrix derived from a global marine food web dataset, resolved to 1° grid cells.
Albouy et al. (2019) — The marine fish food web is globally connectedSpecies occurrences in global freshwater drainage basins. Species names verified against FishBase taxonomy.
Tedesco et al. (2017) — A global database on freshwater fish species occurrence in drainage basinsSpecies distribution range polygons from the IUCN Red List. Freshwater-tolerant species identified via FishBase ecology data ( Fresh flag).
IUCN Red List — Spatial data downloadPrimer pairs
All primer pairs currently included in the database:
| Primer pair | Forward (5’→ 3’) | Reverse (5’→ 3’) |
|---|---|---|
| 12S | ||
| Bylemans |
GCCTATATACCGCCGTCG
|
GTACACTTACCATGTTACGACTT
|
| Evans — ac12s |
ACTGGGATTAGATACCCCACTATG
|
GAGAGTGACGGGCGGTGT
|
| Evans — Am12s |
AGCCACCGCGGTTATACG
|
CAAGTCCTTTGGGTTTTAAGC
|
| Kelly |
ACTGGGATTAGATACCCC
|
TAGAACAGGCTCCTCTAG
|
| Miya — MiFish |
GTCGGTAAAACTCGTGCCAGC
|
CATAGTGGGGTATCTAATCCCAGTTTG
|
| Miya — MiFishE |
GTTGGTAAATCTCGTGCCAGC
|
CATAGTGGGGTATCTAATCCTAGTTTG
|
| Riaz — Vert01 |
TAGAACAGGCTCCTCTAG
|
TTAGATACCCCACTATGC
|
| Taberlet — teleo2 |
AAACTCGTGCCAGCCACC
|
GGGTATCTAATCCCAGTTTG
|
| Taberlet — Elas02 |
GTTGGTHAATCTCGTGCCAGC
|
CATAGTAGGGTATCTAATCCTAGTTTG
|
| Taberlet — Chon01 |
ACACCGCCCGTCACTCTC
|
CATGTTACGACTTGCCTCCTC
|
| Valentini — teleo |
ACACCGCCCGTCACTCT
|
CTTCCGGTACACTTACCATG
|
| 16S | ||
| DiBattista |
GACCCTATGGAGCTTTAGAC
|
CGCTGTTATCCCTADRGTAACT
|
| Evans — Ac16s |
CCTTTTGCATCATGATTTAGC
|
CAGGTGGCTGCTTTTAGGC
|
| Evans — ve16s |
CGAGAAGACCCTATGGAGCTTA
|
AATCGTTGAACAAACGAACC
|
| Kitano |
GCCTGTTTACCAAAAACATCAC
|
CTCCATAGGGTCTTCTCGTCTT
|
| McInnes |
AGCGYAATCACTTGTCTYTTAA
|
CRBGGTCGCCCCAACCRAA
|
| Palumbi |
CGCCTGTTTATCAAAAACAT
|
CCGGTCTGAACTCAGATCACGT
|
| Shaw |
CGAGAAGACCCTWTGGAGCTTIAG
|
GGTCGCCCCAACCRAAG
|
| Cytochrome b | ||
| Kocher |
AAAAACCACCGTTGTTATTCAACTA
|
GCDCCTCARAATGAYATTTGTCCTCA
|
| Miya |
TTCCTAGCCATACAYTAYAC
|
GGTGGCKCCTCAGAAGGACATTTGKCCYCA
|
| Thomsen |
TCCTTTTGAGGCGCTACAGT
|
GGAATGCGAAGAATCGTGTT
|
| COI | ||
| Fields — SharkMiniF1 |
TCAACCAACCACAAAGACATTGGCAC
|
AAGATTACAAAAGCGTGGGC
|
| Fields — SharkMiniF2 |
TCGACTAATCATAAAGATATCGGCAC
|
AAGATTACAAAAGCGTGGGC
|
| West — ElasDegF1 |
ACCAACCACAAAGANATNGGCAC
|
GATTATTACNAAAGCNTGGGC
|
Contribute
GAPeDNA can be extended to additional taxonomic groups. If you have spatialised species checklists and/or metabarcoding primer pairs for a group not yet covered, please get in touch — I can integrate it into the app.
Citation & Credits
If you use GAPeDNA in your work, please cite:
Diversity and Distributions, 2021
Development & maintenance: Virginie Marques
Illustration: P. Lopez
Server infrastructure: M. Q. Quidoz (CEFE)